If you have used the Gedmatch One-to-One Comparison tool, then you have seen total half-match segments (HIR). In this post, find out what this means and how to apply it to your genetic genealogy research.
As many users of Gedmatch will tell you, most of their shared segments with their matches are half-match regions. Sharing half-match regions or segments with your DNA relatives is normal, and you will learn why that is in this article.

Once you understand the how and why behind the half-match segments, you will be well on your way to understanding more about your DNA matches and how to use them to learn about your family tree and ancestry.
What is a Half-Match segment or HIR on Gedmatch?
A half-match segment (HIR) is a DNA segment in a half-identical region of shared DNA between you and a DNA match. It is half-identical because it is only identical on one copy of your chromosome at that location.
In the image below from Gedmatch One-to-One Comparison Tool results, you can see an example of half-identical DNA segments shared between half-siblings. We note that these half-sisters share 1627.8 centimorgans of half-match segments.
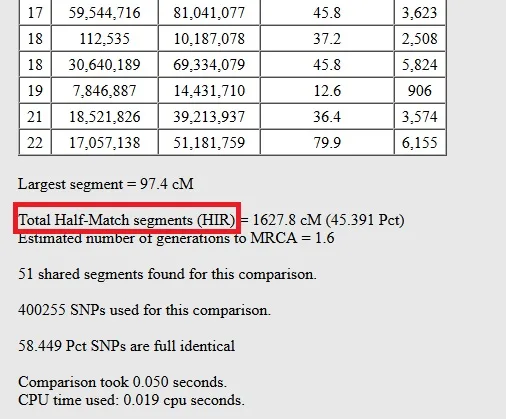
So, in this context, what does "HIR" stand for? It's the half-identical region that I mentioned earlier.
In the table below, we can see the chromosome number, and the start and end location for each half-matching DNA segment. The third column is the genetic distance of the length of the segment, measured in centimorgans.
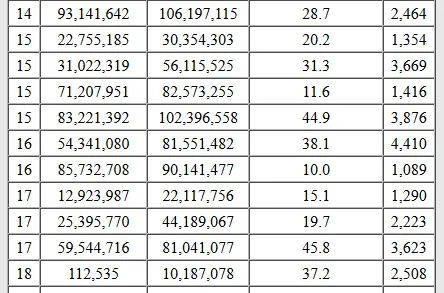
Aren't all DNA segments half-identical? There are situations where a DNA segment is identical on both copies of the same chromosome at the same location.
What does it mean to share half-identical regions?
Two people who share half-identical regions inherited DNA segments that are identical to the other person from one side of their family.
It does not automatically mean that you and your DNA match have a half-relationship, necessarily.
Half-match DNA segments are the most common type of DNA segments. We will share these types of segments with our full-siblings, half-siblings, our parents, as well as close and distant cousins.
Take, for example, the half-sisters that I mentioned above. Their DNA segments are half-identical because they inherited their identical DNA regions from their common mother.
This is because, generally, we are related to most of our DNA matches on only one side of our family tree. Exceptions to this would include close relatives, such as siblings, or distant relatives who are related on both sides of our family due to endogamy.
Below, you can see another example from two people who are second cousins once-removed. The common ancestor, the great-grandparent of the oldest of the two test takers, passed down the following segments through the paternal line of the younger DNA match.
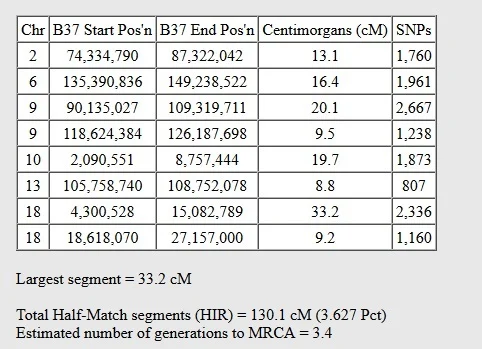
Because the younger match is only related to this DNA match on her father's side of the family, there are no fully-identical DNA regions. In order for her to have inherited fully identical regions, she would have had to inherit identical segments from both her mother and father.
What does each half-match region on Gedmatch represent?
As you might have guessed, each half-match region over a particular size represents a DNA segment passed down from a common ancestor. You may have inherited more than one DNA segment from the same ancestor long ago.
Most experts recommend disregarding half-match segments that are smaller than 7 centimorgans in length. These small segments can be real, but they are more often false.
The reason behind this is because the smaller the DNA segment, the higher the probability that the shared DNA segment is coincidentally identical. DNA segments that are coincidentally identical were not inherited from a shared ancestor.
It is possible to have multiple common ancestors shared with the same DNA match, with each half-match region representing those multiple shared lines of ancestry.
The best way to determine which shared ancestor passed down your half-match segments is to compare your family tree with that of your DNA match.
The following graphic is a visual representation of half-identical regions, which are also commonly referred to as half-identical segments. The black arrows point to (simplified) DNA segments located on one copy of each chromosome that are identical at the location.
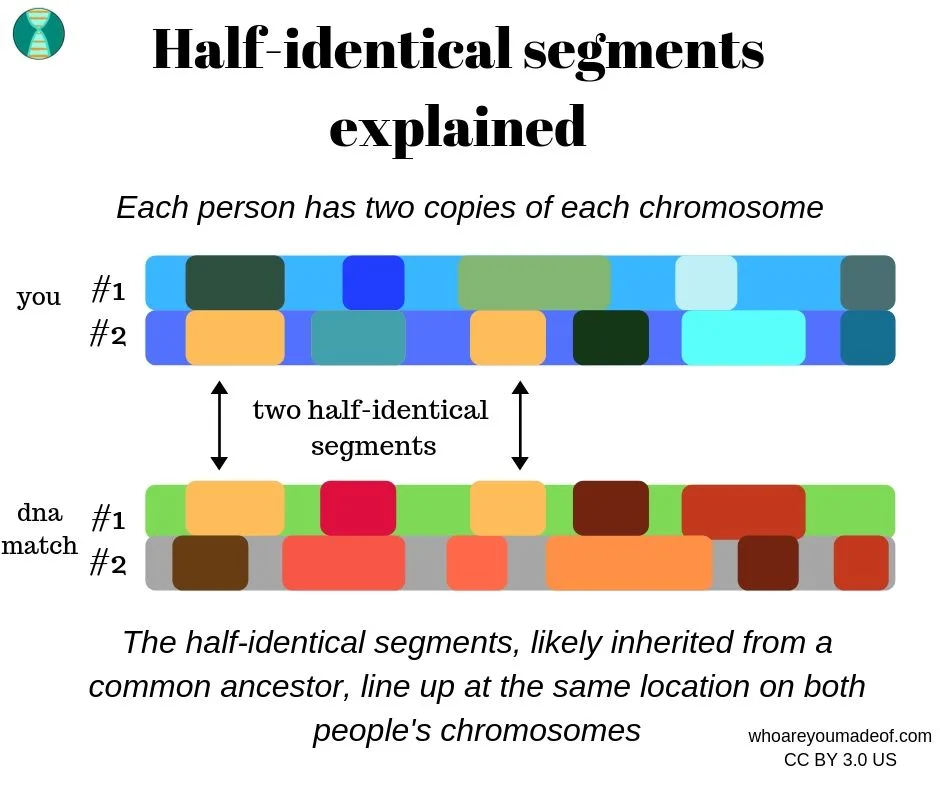
As you may have noticed in the illustration above, each DNA tester has half-identical DNA segments on a different copy of each chromosome, but at the same location. This is normal, and the only thing that we need to know is that we share identical segment(s) at the same location on at least one copy of a chromosome.
These segments are half-matching because they only match on one copy of the chromosome and not on both copies. If they matched on both copies, which is possible in close relatives, such as full-siblings, we would label them fully-identical identical regions or segments.
Conclusion
I hope that this post helped you learn more about half-match segments on Gedmatch, what they mean, and what each region represents.
If you have any questions about something that you read in this post, or if you would like to share your own experience working with these regions, please join us in the discussion below.
Thanks for reading 🙂



Pamela Silva
Sunday 2nd of July 2023
I could really use some advice or input. I wrote once before but I did not have enough understanding of DNA or reading matches.
My closest match is someone I did not know. A female. We both did the DNA test years ago and have been friends ever since. She was looking for her biological father. I was finishing a promise to my late mom to discover her side of the family. Well, this year I decided I needed to finish what I started. I found your newsletter, which has been a tremendous help. I also have uploaded both DNA samples to Gedmatch. I have tried to learn as much as I could. At any rate, the question still remains, are we or are we not, half-sisters? Ancestry's Match says this:
Shared DNA: 1,688 cM across 41 segments Unweighted shared DNA: 1,688 cm Longest segment: 122 cm
BUT Gedmatch, when I run the one-to-many shows the results.-105.4 1715.3 1.5 274123 CM's 1715.3 105 longest segment and we are only 1.5 generations away.
Then the one to one X =results are the following;
Chr B37 Start Pos'n B37 End Pos'n Centimorgans (cM) SNPs Segment threshold Bunch limit SNP Density Ratio 23 2,700,157 52,067,217 82.7 9,093 186 111 0.65 23 61,938,671 154,929,412 105.1 12,553 188 112 0.63
Largest segment = 105.1 cm
Total segments 187.9cM (98.898 Pct)
2 shared segments were found for this comparison.
22003 SNPs were used for this comparison.
Since GEDMATCH has added a space on Relationship Probability in which Briton says makes the result more accurate, which I just had to give up. I have done both our trees. Our families are very integrated. Along with one other family. And adding more confusion, a new "cousin" match is now showing as our half-brother. If I could get some help from anyone to pull myself out of this mire, I would be so very appreciative.
Virginia R
Saturday 27th of April 2024
@Pamela Silva, this is very interesting. Has anyone replied with an answer?