Have you noticed that shared DNA segments with the same relative on different testing sites don't have identical locations on the chromosome? In this post, find out the reason why this happens.

One of the most helpful aspects of more advanced DNA analysis is the ability to locate specific DNA segments shared with several matches in order to identify the common ancestor. Sometimes all of our DNA matches are not located on the same site, and we find ourselves comparing chromosome locations across DNA testing platforms and third-party websites.
Recently, a very good question was raised: Is the information about start and end position of a DNA segment the same, no matter which testing company or website provides you the data?
In this post, we'll dive into chromosome positions across testing sites.
The short answer to this question is that the start and end positions are generally similar, but not identical. In order to demonstrate this, I'll show you comparisons between my own DNA and that of my relative, and you'll see start and end positions from Gedmatch, Family Tree DNA, and My Heritage DNA.
You'll be able to draw your own conclusions about how to proceed in interpreting matching segments with your own DNA matches.
Example of chromosome positions across testing sites
I have a fairly close relative who initially tested with Family Tree DNA, but has uploaded their DNA to Gedmatch, and My Heritage DNA. I tested with Ancestry DNA initially, but have uploaded to all of the same sites that he did, as well as Family Tree DNA.
In the image below, you can see a table comparing three distinct DNA segments across all four of these websites.
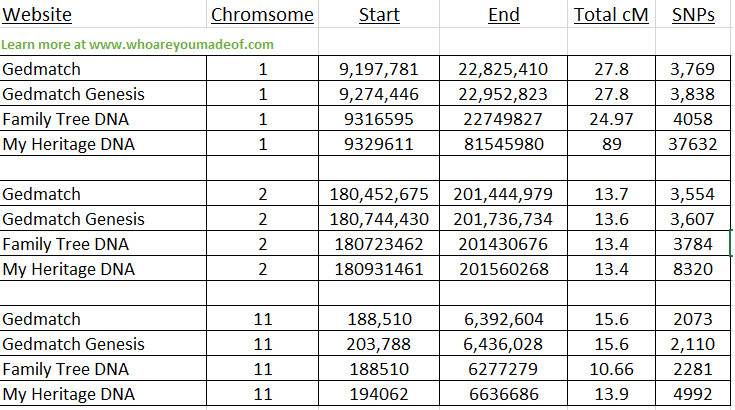
As you can see, the start and end positions are generally very close to each other. There are slight differences because of the methodology used by each site and the fact that the kits were tested by different companies - meaning there is slight variation in the data used to make the comparisons.
If you noticed that there is a big difference in one My Heritage segment, it's not your imagination. You can read about this below.
Segment Stitching at My Heritage
My Heritage "stitches" together adjacent segments. If you look at the image below, you can see that the start positions of the first matching segment on the first chromosome are very similar, but that the Family Tree DNA segment ends, and the My Heritage DNA segment keeps going - for much longer.
This is because I share two segments with this relative on Chromosome 1, and it happens to be very close to the first one.
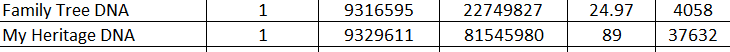
Just to get an idea of how close the chromosomes are, you can take a peek at the location of the two segments, per Family Tree DNA:

The stitching of the two very close DNA segments doesn't have a major effect on the end result of the comparison: this relative is fairly closely related to me. In fact, since this relative did not test their DNA with the same company, this segment might actually be continuous, and My Heritage just put it back together.
Every company "reads" different SNPs, and there is a chance that My Heritage inferred the data that was not included in our test results (especially since neither of us testing with My Heritage DNA).
DNA matches, overlapping segments, and start and end positions
When it comes to the differences between start and end positions across testing companies and platforms, you might wonder if the different start and end positions will affect your genealogy research.
It can be helpful to "triangulate" a matching segment with a few different matches to determine a common ancestor, and this is the main reason that genetic genealogists (such as yourself) would be concerned about these discrepancies.
Do differences in start and end positions mean anything?
You can see from the table that I posted at the beginning of the post that the segments are relatively the same size, so as long as the start and end positions are similar, there is no real consequence to them having slightly different start and end positions.
If I had other relatives that overlapped on these segments, I could possibly conclude that they are descended from the same common ancestors that I share with this particular close relative.
On the note of shared segments and common ancestors, there is one more important issue to mention.
Shared segments don't always mean a shared ancestor among matches
Wait, what? Just when we really feel like we've gotten the hang of this genetic genealogy business, we learn that shared segments among matches don't always mean that we all share the same common ancestor.
What do I mean by this? Imagine that you have three DNA matches that all have segments that match you, in about the same range (they'll never be exact) on Chromosome 1.
It's easy to assume that all four of you (you, plus your three matches) all inherited this segment from the same common ancestor.
Two copies of each chromosome
Since we inherit a copy of each chromosome from each of our parents, it is always possible that the segment that you share with one or more of your matches could have been inherited from the other site of your family. This is really good to keep in mind - if you are searching through family trees looking for common surnames, it could be that you are barking up the wrong tree - almost literally.
In our example of the three DNA matches that have overlapping segments on Chromosome 1, there is a good possibility that you share common ancestors with all of them, but there is a chance that they don't all share common ancestors with each other.
You might share an ancestor from your mom's side of the family with two of them, and an ancestor from your dad's side of the family with the other one.
Ignore really small segments
I don't want to go too deep into details about segment size, but it's always good to keep in mind that the smaller the segment, the bigger the chance that it is just coincidentally identical and does not indicate a common ancestor.
It's good practice to completely ignore segments small than 5 centimorgans (cMs), and to view segments smaller than about 10 cMs with a healthy dose of skepticism. I prefer to only focus on segments larger than 15 cMs for serious research, since it takes a lot of time to research a DNA match.
Conclusion
I hope that this post has helped you see the minor differences between start and end positions on segments that you share with matches across the testing sites, and how these differences could lead to confusion if misinterpreted.
If you have any questions about something that you read in this post, I encourage you to leave a comment below. I hope to hear from you.
Thanks for stopping by!


Gary Kmosko
Friday 5th of August 2022
Is there a way to distinguish if a segment match is maternal or paternal? Or, when an overlapping segment match switches from maternal to paternal?